Which Of The Following Chromatin Changes Will Lead To Upregulation Of Expression?
- Research
- Open Admission
- Published:
Leo1 is essential for the dynamic regulation of heterochromatin and gene expression during cellular quiescence
Epigenetics & Chromatin volume 12, Article number:45 (2019) Cite this commodity
Abstruse
Groundwork
Cellular quiescence is a reversible differentiation state during which cells modify their cistron expression plan to inhibit metabolic functions and arrange to a new cellular environment. The epigenetic changes accompanying these alterations are not well understood. Nosotros used fission yeast cells as a model to study the regulation of quiescence. When these cells are starved for nitrogen, the cell bike is arrested in G1, and the cells enter quiescence (G0). A factor regulatory program is initiated, including downregulation of thousands of genes—for example, those related to cell proliferation—and upregulation of specific genes—for case, autophagy genes—needed to conform to the physiological challenge. These changes in factor expression are accompanied by a marked alteration of nuclear system and chromatin construction.
Results
Here, we investigated the office of Leo1, a subunit of the conserved RNA polymerase-associated factor one (Paf1) complex, in the quiescence process using fission yeast every bit the model organism. Heterochromatic regions became very dynamic in fission yeast in G0 during nitrogen starvation. The reduction of heterochromatin in early G0 was correlated with reduced target of rapamycin complex two (TORC2) signaling. We demonstrated that cells lacking Leo1 show reduced survival in G0. In these cells, heterochromatic regions, including subtelomeres, were stabilized, and the expression of many genes, including membrane transport genes, was abrogated. TOR inhibition mimics the effect of nitrogen starvation, leading to the expression of subtelomeric genes, and this effect was suppressed by genetic deletion of leo1.
Conclusions
We identified a protein, Leo1, necessary for survival during quiescence. Leo1 is part of a conserved protein complex, Paf1C, linked to RNA polymerase II. We showed that Leo1, acting downstream of TOR, is crucial for the dynamic reorganization of chromosomes and the regulation of gene expression during cellular quiescence. Genes encoding membrane transporters are non expressed in quiescent leo1 mutant cells, and cells die afterward 2 weeks of nitrogen starvation. Taken together, our results suggest that Leo1 is essential for the dynamic regulation of heterochromatin and gene expression during cellular quiescence.
Background
When fission yeast cells are starved for nitrogen, they quickly divide twice without growth, arrest in the G1 phase of the cell wheel and enter cellular quiescence (G0). A gene regulatory program is initiated, including downregulation of thousands of genes—for example, genes related to cell proliferation—and upregulation of specific genes, such as autophagy genes, necessary for adaptation to the physiological challenge [1,2,three]. Thus, the transcriptional profile of quiescent S. pombe cells is dramatically reprogrammed to allow survival under low-nitrogen conditions [4]. This change is accompanied past a marked alteration of nuclear organization. The cell bike arrest is reversible, and when nitrogen is added, quiescent cells readily exit the arrest and begin proliferating. Recently, two studies demonstrated that the RNA interference pathway is essential for viability during cellular quiescence. Dicer is of import for heterochromatin assembly at centromeres during entry into G0 and for preventing heterochromatin formation over rDNA repeats in G0 cells [5]. Argonaute (Ago1), together with the H3K9 methyltransferase Clr4, plays a function in the repression of euchromatic genes in quiescent cells [6]. Together, the results of these studies clearly testify that dynamic regulation of heterochromatin is essential during quiescence in fission yeast.
Leo1 is a subunit of the RNA polymerase-associated factor 1 (Paf1) circuitous (Paf1C). Paf1C consists of 5–half dozen subunits with four core subunits, including Leo1, which are conserved in eukaryotes. The complex has been implicated in several functions linked to RNA Pol Two, including transcription elongation and mRNA 3' stop germination, and interacts with histone modifying enzymes, transcriptional activators, elongation factors and RNA cleavage factors [7]. A function for Paf1 in cell proliferation and last differentiation during development has been described in Drosophila [8]. Paf1C is also implicated in epigenetic regulation. For example, mutations in genes encoding Paf1 subunits raise heterochromatin silencing past the RNAi pathway [9]. We previously institute that two of the Paf1C core subunits, Leo1 and Paf1, are important for histone turnover and preclude heterochromatic regions from expanding in vegetative cells [10]. In this study, we addressed the role of Leo1 during cellular quiescence.
Results
Leo1 is required for survival during long-term quiescence
To identify additional factors involved in chromatin regulation during cellular quiescence, we applied a pocket-size-scale genetic screen using a selection of mutants affecting chromatin dynamics (Additional file 1: Fig. S1a). Of the twelve candidates tested, only leo1∆ cells showed strongly reduced viability after long-term nitrogen starvation. Subsequently nitrogen starvation, leo1∆ cells entered quiescence with kinetics similar to those of wild-type (WT) command cells (Fig. 1a, b); however, these cells exhibited a phenotype of reduced survival in G0. While the wild-type strain maintained a survival rate of 99.2% afterward 21 days in G0, the survival rate of leo1∆ cells remained 99.4% at 7 days and then gradually decreased to 49.iv% at 21 days (Fig. 1c). For comparison, the clr4∆ cell survival rate was also decreased, in agreement with a previous written report [6]. To investigate whether the other components of the Paf1C complex are too important during quiescence, we measured the survival rate of strains carrying genetic deletion of the Paf1C core components leo1, paf1, cdc73, and tpr1 (ctr9) during a 3-week quiescence time course experiment (Additional file i: Fig. S1b, c). We also included the prf1 mutant, but it was omitted from the survival assay due to its peculiar cell morphology. The different mutations resulted in reduced survival of the host cells later 2–3 weeks of nitrogen starvation. Thus, Leo1 and three other cadre components of the Paf1C circuitous are required for survival during long-term quiescence.
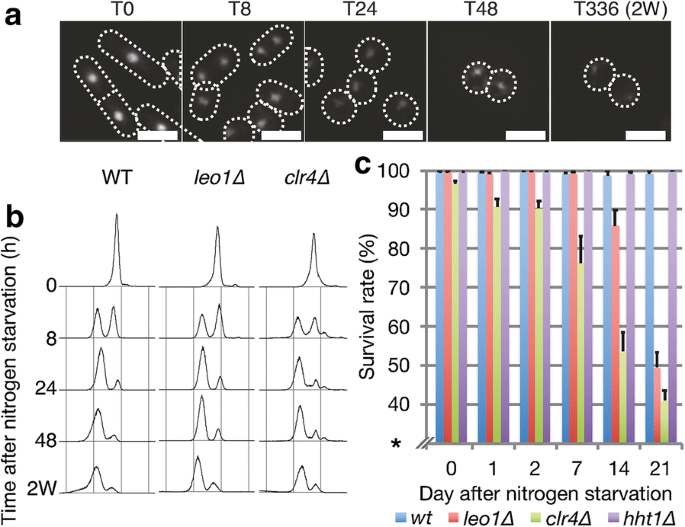
Leo1 is required for survival during long-term quiescence. a DAPI-stained images of wild-type cells at various time points during nitrogen starvation. Scale confined: ten µm. b DNA content assay by flow cytometry at various time points during nitrogen starvation. c Survival charge per unit of the indicated strains at various time points during nitrogen starvation. The mistake confined represent the standard deviations (SDs) of three biological replicates
Leo1 affects the dynamics of heterochromatin assembly/disassembly at constitutive heterochromatic regions
We hypothesized that uncontrolled heterochromatin associates in leo1∆ cells limits the lifespan of these cells during long-term quiescence. To investigate this possibility, we performed genome-wide mapping of histone H3 di-methylation (H3K9me2) in wild-blazon and leo1∆ cells during nitrogen starvation. Interestingly, in wild-type cells, the H3K9me2 levels in all heterochromatic regions that are constitutive in vegetative cells, i.east., centromeres, the mating-type region and telomere-proximal regions, diminished in early G0 phase (later viii and 24 h) and were and so partially restored afterwards 48 h (run into Fig. 2b, tel1L:0–20 kb; Boosted file 1: Fig. S1). In sharp contrast, the H3K9me2 levels remained constant in leo1∆ cells throughout G0 phase (Fig. 2; Additional file 2: Fig. S2). In subtelomeric regions of chromosomes I and II, the H3K9me2 levels in wild-blazon cells varied depending on the distance from the telomere and the time point during G0. In telomere-proximal regions (0–twenty kb; 20–xl kb) wild-type H3K9me2 levels decreased at 8 and 24 h and so increased again at 48 h. In contrast, the subtelomeric H3K9me2 levels in leo1∆ cells remained high throughout G0 phase (Fig. 2b; Additional file ii: Fig. S2). The wild-type H3K9me2 levels in pericentromeric regions (imr and otr) too varied depending on the fourth dimension point in G0, whereas these levels in leo1∆ cells were much less dynamic (Fig. 2c; Additional file 2: Fig. S3). Finally, the same trends were observed in the mating-type region (mat) (Fig. 2d).
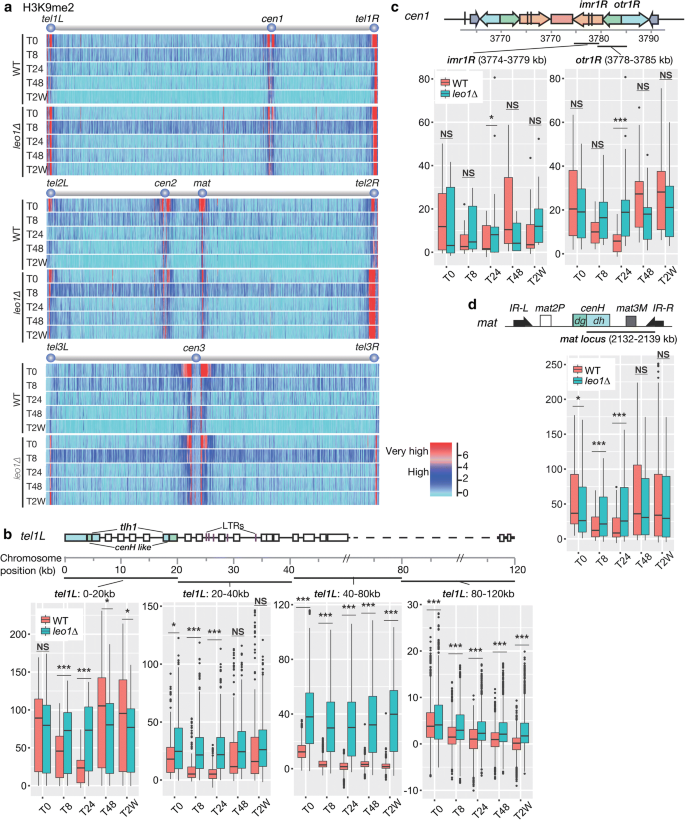
Loss of Leo1 causes mis-regulation of heterochromatin. a Mapping of H3K9 methylation in wild-type and leo1∆ cells. The relative fold enrichment of di-methylated H3K9 (log2), every bit adamant by ChIP-scrap analysis with two biological replicates, is plotted. b–d Box plots showing changes in Scrap-chip signals for H3K9me2 at constitutive heterochromatin loci: the subtelomeric region tel1L (b); the pericentromeric region cen1; and the mating-blazon locus (d). The eye lines evidence the medians; the box limits indicate the 25th and 75th percentiles as determined by R software; the whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles, and outliers are shown. The y-axes show the relative enrichment of H3K9me2. The p values are from contained-samples t-test comparison the hateful values between leo1∆ and WT at unlike fourth dimension points. (NS) Non-meaning; (*) p value < 0.05; (**) p value < 0.01; (***) p value < 0.001
To examination if the conversion of H3K9me2 to me3 is afflicted by leo1Δ cells we performed Flake-QPCR for H3K9me3 at several heterochromatic loci at T0, T24 and T48 during quiescence (Additional file 3: Fig. S10). Nosotros find reduced H3K9m3e levels at all tested loci in wild-type cells at T24 and T48 compared to T0. This shows that the reduced H3K9me2 levels in quiescent cells are non due to conversion to H3K9me3. We also observe increased H3K9me3 levels in leo1Δ cells compared to wild type at all heterochromatic loci, peculiarly at subtelomeric genes SPAC186.04c and SPBPB2B2.18 (Additional file 3: Fig. S10).
These observations show that constitutive heterochromatic regions in vegetative cells become subject to dynamic regulation during cellular quiescence leading to a transient reduction of both H3K9me2 and me3 and that Leo1 is required for this dynamic behavior of heterochromatin.
Leo1 is disquisitional for proper heterochromatin associates at heterochromatin islands during G0 entry
In improver to subtelomeric heterochromatic regions, other H3K9me2-enriched regions were found in leo1∆ cells relative to the H3K9me2 enrichment patterns in wild-type control cells (Additional file 2: Fig. S4). These regions overlapped with the previously identified islands of facultative heterochromatin, for example, at the mcp7 +, mei4 +, and ssm4 + loci [11]. The genes in these islands are regulated past the RNA elimination mechanism (DSR) and the RNA processing complex (MTREC) [12]. In leo1∆ cells, the H3K9me2 levels at these islands were approximately twofold higher than those in wild-type vegetative cells, decreasing gradually in early G0 and further diminishing after 2 weeks. These data advise that Leo1 is critical for proper heterochromatin dynamics at DSR/MTREC-dependent heterochromatin islands during entry into G0 phase.
Loss of Leo1 leads to the repression of genes located near the telomeres of chromosomes I and II
Genome-wide mapping of H3K9me2 showed that Leo1 counteracts heterochromatin assembly, which is normally associated with transcriptional repression. To determine whether Leo1 affects gene expression in G0, we performed RNA sequencing (RNA-seq) assay in wild-blazon and leo1∆ cells after the shift to nitrogen-costless medium. As expected, the expression of many genes was lower in leo1∆ mutant cells than in wild-type cells, peculiarly at 24 and 48 h after the shift (Fig. 3a). When plotted forth the chromosomes, the downregulated genes were enriched virtually the telomeres of chromosomes I and II (Fig. 3b and Additional file 2: Fig. S5). In contrast, the leo1∆ mutation did not affect gene expression at the ends of chromosome III, which containing rDNA repeats. To investigate the statistical significance of telomere proximity for the genes downregulated in leo1∆ versus wild-type cells, we performed a Chi-foursquare examination (Additional file ii: Fig. S8), which revealed that the downregulated genes were significantly clustered in subtelomeric regions 0–eighty kb from the telomere at tel1L, tel1R, tel2L and tel2R. This effect was observed starting viii h subsequently nitrogen removal and persisted throughout the 2-calendar week quiescence time form. Thus, these results clearly evidence that loss of Leo1 leads to the repression of genes throughout the genome, with a stronger effect for genes nigh the telomeres of chromosomes I and Two.
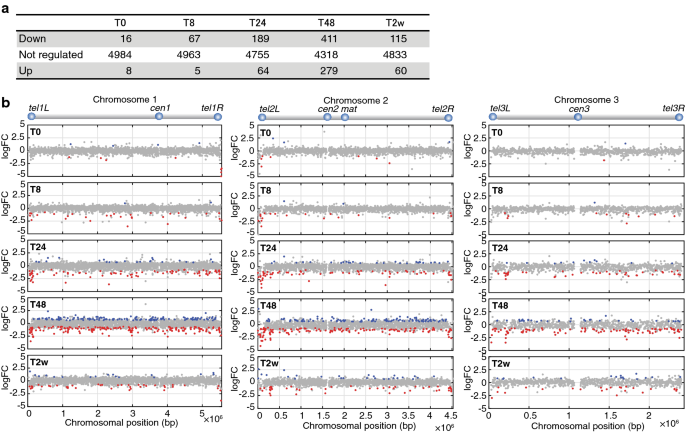
Loss of Leo1 leads to repressed transcription of genes located near the telomeres of chromosomes I and 2. a The tabular array shows the numbers of differentially regulated genes between leo1∆ and wild-type cells at each time bespeak during quiescence. The R/Bioconductor package EdgeR was used for the assay of differential gene expression. For a cistron to be deemed meaning, the FDR adjusted p value had to be less than 0.05. 'Down', 'Non inverse' and 'Up' bespeak genes downregulated, unchanged and upregulated, respectively, in leo1∆ cells relative to their expression in wild-type cells. Genes were filtered to remove genes with very low or no expression from the assay. b Genome-wide distribution of regulated genes in leo1∆ cells and wild-blazon cells. The x-axis indicates the chromosomal positions (× 10−vi bp), and the y-axis shows the log2 fold alter (log2FC) values for genes differentially expressed betwixt leo1∆ and wild-type cells. Up- and downregulated genes are shown in blue and ruby, respectively (FDR < 0.05)
Loss of Leo1 prevents the expression of membrane transporter-encoding genes in G0
Next, we analyzed the expression patterns of all differentially expressed genes using hierarchical clustering. The differentially expressed genes clustered into ix major clusters (cl) based on the patterns of expression at different time points. The clusters are shown on a rut map and in a graphical format (Fig. 4a, b). Four clusters exhibited lower expression in leo1∆ cells than in wild-type cells, with decreasing (cl1) or increasing (cl2, iii and four) expression levels during early on G0. Three clusters exhibited college expression in leo1∆ cells, with increasing (cl5 and 7) or decreasing (cl6) expression levels during early on G0. The terminal two clusters were mirror images, with a peak at T8 (cl8) or T48 (cl9).
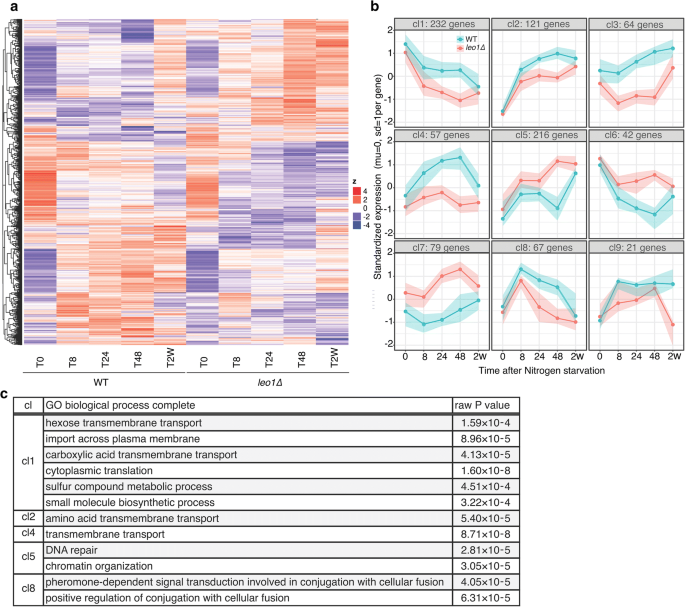
Loss of Leo1 causes the downregulation of membrane transporter-encoding genes. a A heatmap of all the genes found to be significant in whatsoever of the comparisons shown in Fig. 3. The expression of each gene was standardized (mean = 0, SD = 1), and the genes were then clustered by hierarchical clustering. b The clustering performed for the heatmap was limited to nine clusters and plotted as lines to enhance the visual representation of the gene regulation patterns. The lines bespeak the medians of the cluster, and the colored areas surrounding the lines show the 25th and 75th percentiles, significant that l% of the genes were in the cluster. c List of the Get biological process terms enriched with the differentially expressed genes in the clusters. The significance of the Become terms was determined using the FDR correction (p< 0.05). The granular terms with the largest fold enrichment are shown for each group. Clusters cl3, cl6, cl7 and cl9 had no apparent Go term enrichment
To gain insight into the biological processes regulated during quiescence, we performed Factor Ontology (GO) analysis of differentially expressed genes within these nine clusters and identified the associated enriched Become terms (Fig. 4c). In clusters 1, 2, and 4, which showed lower expression in leo1∆ cells than in wild-type cells during quiescence, Go terms for the processes of trans-membrane transport of hexose and amino acids were significantly enriched (FDR < 0.01). In cluster 5, which showed higher expression in leo1∆ cells during quiescence, the Become terms for the processes of Deoxyribonucleic acid repair and chromatin organization were significantly enriched. Finally, for cluster 8, with a peak at T8, the Get term for conjugation with cellular fusion was significantly enriched.
Paf1C stimulates heterochromatic histone turnover in G0 cells
We previously showed that the turnover of histone H3 in heterochromatic regions of fission yeast is reduced in vegetative cells lacking the Leo1 subunit of Paf1C [10]. To investigate whether histone turnover was affected in G0 cells, we performed recombination-induced tag substitution for histone H3 (H3-RITE) experiments (Fig. 5a). For this analysis, we focused on a prepare of four subtelomeric genes repressed in vegetative cells by Paf1C [13] (Fig. 5b). Starting time, we measured the recombination rates by scoring the frequency of hygromycin-sensitive colonies. At T0, later on ii h of Cre recombinase induction, wild-type and leo1∆ cells exhibited recombination rates of 48.1% and 49.four%, respectively. At T24, subsequently 4 h of Cre recombinase induction, wild-type and leo1∆ cells exhibited recombination rates of 35.4% and 35.5%, respectively. Next, we measured histone commutation but institute no meaning increase in T7-tagged histones after tag substitution, suggesting that the incorporation of new histones is very slow in quiescent cells. Withal, we did find reduced levels of old (HA-tagged) histones in wild-blazon cells afterward 24 h in G0 (Fig. 5c). This reduction was dependent on Leo1, since the level of old histones was not significantly reduced in leo1∆ cells. Thus, we concluded that Paf1C stimulates the eviction of old histones in G0 cells.
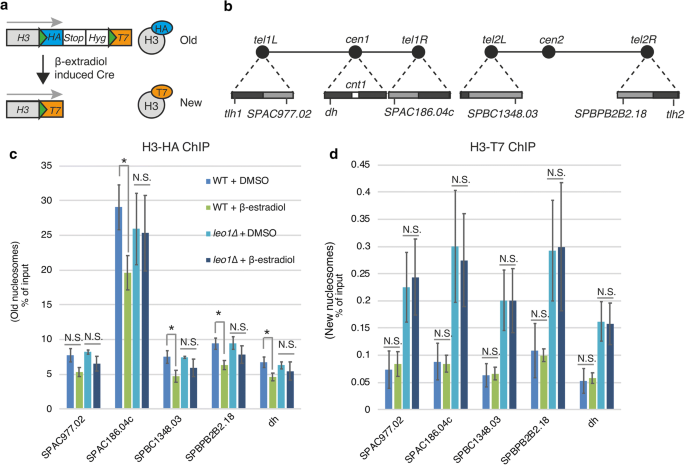
Leo1 regulates histone eviction during quiescence. a Diagram of the RITE system. The green triangles represent LoxP sites. b A schematic of fission yeast chromosomes I and Ii. The triangles on the chromosomal edges are enlarged views of centromeric and subtelomeric regions. The nighttime and calorie-free grey boxes indicate regions of loftier and low H3K9me2, respectively. Scrap of H3-HA (c) and T7-HA (d) in silent chromatin regions. The values are the averages of three biological replicates. The error bars point SDs; p values (paired Student's t examination): *< 0.05; Northward.South. (not meaning) > 0.05
The activity of the TORC2/Gad8 pathway is modulated during quiescence
To investigate the means by which Leo1 regulates heterochromatin during quiescence, we focused on target of rapamycin (TOR) circuitous 2 (TORC2). TORC2 responds to glucose levels, and considering it activates the protein kinase Gad8 (an orthologue of human AKT), information technology is required for the regulation of prison cell cycle progression, starvation responses, and cell survival [14,15,sixteen]. In add-on to these functions, TORC2 was recently reported to be required for gene silencing and the formation of heterochromatin in fission yeast. Tor1 and Gad8 are needed for the silencing of subtelomeric heterochromatic regions [13]. The function of Tor1/Gad8 is linked to Paf1C, since the silencing defects in subtelomeric regions in tor1∆ and gad8∆ mutant cells are fully suppressed by genetic deletion of paf1 [13]. Based on this finding, nosotros hypothesized that a decrease in TORC2 activeness could induce the transient reduction in H3K9me2 levels at subtelomeric regions at 24 h after nitrogen starvation. Therefore, nosotros monitored the protein level of activated Gad8 during quiescence. Western blotting was carried out using specific antibodies against the active phosphorylated form of the Gad8 kinase (Fig. 6a; Boosted file two: Fig. S9). Although some variation between biological duplicates was observed, all results showed a similar tendency: the level of activated Gad8 clearly macerated in early G0 phase (after 8 and 24 h) and was subsequently restored afterwards 48 h and 2 weeks. These changes in Gad8 activity were inversely correlated with the observed Leo1-dependent changes in H3K9me2 in heterochromatic regions during G0 phase (Fig. 2; Additional file 2: Figs. S2, S3). Moreover, we found no change in Gad8 activation between leo1∆ cells and wild-type cells (Boosted file 2: Fig. S9), suggesting that Gad8 acts upstream of Paf1C.
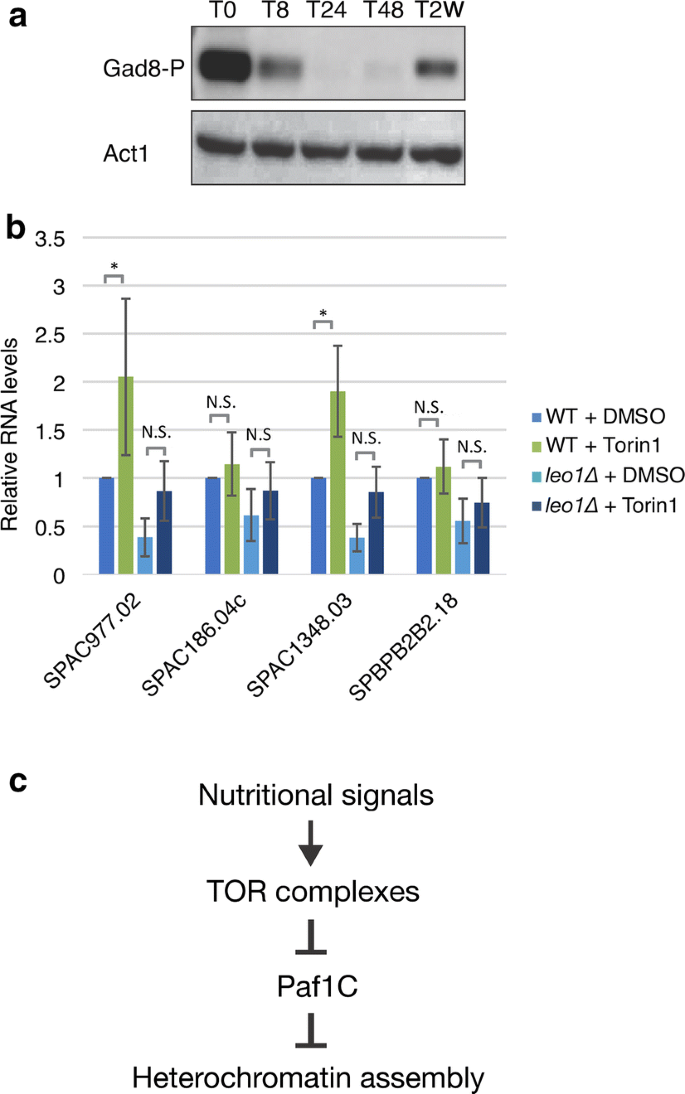
The Tor1/Gad8 pathway regulates Paf1C. a Phosphorylation of Gad8 at diverse time points after nitrogen starvation. b RNA expression levels of subtelomeric genes on chromosome I and II were quantified by RT-qPCR. Cells were treated with DMSO or 25 µM Torin1. Human being RNA was used every bit the fasten-in control RNA for normalization of the expression levels. The values are the averages of three biological replicates (n = 6). The error bars indicate SDs; p values (paired Student's t exam): *< 0.05; N.S. (not meaning) > 0.05. c A proposed model for the regulation of heterochromatin via TOR complexes and Paf1C in G0 cells (see the main text for details)
To examine the effect of TOR activeness on the expression of subtelomeric genes, we used the TOR inhibitor Torin1 and studied cistron expression by RT-PCR using the aforementioned fix of four genes (Fig. 6b). Torin1 treatment clearly mimicked nitrogen starvation conditions, leading to a significant increase in the expression of these genes (Fig. 6b). However, handling of leo1∆ cells with Torin1 did non atomic number 82 to significant upregulation (Fig. 6b). Therefore, we advise that Paf1C activeness in G0 cells is dependent on TOR complexes (Fig. 6c).
Give-and-take
A machinery of chromatin dynamics involving the Leo1 subunit
Interestingly, the construction of Paf1C in circuitous with Pol II reveals that the Paf1 and Leo1 subunits are positioned in front of the Pol II complex near the downstream DNA in a perfect position to meet the incoming nucleosomes during elongation [17]. Therefore, this domain of Paf1C is likely positioned appropriately for interest in nucleosome disassembly during transcription, assuasive the turnover of histones. Hither, we observed reduced levels of one-time histones in G0 cells, and this reduction was dependent on Leo1.
Previous work showed that the turnover of histone H3 in heterochromatic regions of vegetative cells is also dependent on Leo1 [10]. H3K9me2 regions accept recently been shown to be permissive for transcription in fission yeast [xviii]. It is plausible that the increased stability of heterochromatin in quiescent leo1Δ cells is caused by a failure to remove old histone H3, thereby stabilizing the H3K9me2 level. We previously showed that Leo1 counteracts H3K9me2 in vegetative cells [10]. Here, nosotros show that some heterochromatic regions are destabilized in wild-blazon cells and that H3K9me2 levels are reduced during quiescence in a Leo1-dependent manner. Deletion of the leo1 cistron leads to both up- and downregulation of genes relative to their wild-blazon levels during quiescence (Fig. 3a). To farther explore the effects of leo1 gene deletion on gene expression, we directly compared changes in RNA expression levels with changes in H3K9me2 levels (Additional file 2: Figs. S6, S7). This analysis revealed a propensity for downregulated genes to be located in regions with increased H3K9me2 levels in leo1∆ cells, suggesting that direct stabilization of heterochromatin leads to factor repression. Quiescent cells are arrested in the G0 phase of the prison cell wheel and do non undergo Dna replication, which, in vegetative cells, contributes to histone turnover at the replication fork. Therefore, Paf1C action may be peculiarly important in G0 cells to remove old histones marked by H3K9me2 in order to permit epigenetic reprogramming to support the cistron expression changes needed for adaptation to the low-nitrogen surroundings.
Changes in subtelomeric gene expression mediated by Paf1C
We observed decreased expression of genes involved in the transport of hexose and amino acids in leo1∆ cells during G0. This subtract was accompanied by an expansion of heterochromatin in the subtelomeric regions where many of these genes are located. These findings strongly suggest that Paf1C has a special function in subtelomeric heterochromatic regions to permit the reduction of heterochromatin and the induction of gene expression in these regions to adapt to cellular quiescence. Genes in the subtelomeric regions showed Leo1-dependent histone turnover (Fig. 5c) and had lower expression levels in leo1∆ cells than in wild-blazon cells (Fig. 6b). Assay of the time course data showed that downregulation started viii h subsequently the nitrogen shift and that the genes remained repressed throughout the ii-week time course (Boosted file ii: Fig. S8). Since several of the subtelomeric genes encode transporters, the reduced viability of leo1∆ cells observed after 2–3 weeks in G0 may be caused by transport deficiencies due to failure to limited these genes. The phenotypic lag menstruum between the reduced expression and loss of viability in G0 may be due to reductions in the levels of nutrients that become critical after 2 weeks.
Is the function of Paf1 in epigenome dynamics conserved?
A office for the Paf1C component Ctr9 in jail cell differentiation has recently been reported in Drosophila [8]. Drosophila neurons fail to terminally differentiate, and hundreds of genes, including several neuropeptide-encoding genes, are downregulated in ctr9 mutants. A reduction in the activating histone marking H3K4me3 was observed in ctr9 mutants [8]. However, whether H3K9 methylation is increased in Drosophila ctr9 mutants in the same way as in leo1 mutant fission yeast is unknown, and whether Paf1C is important for the heterochromatin changes that occur during cell differentiation and quiescence in other species remains to exist investigated.
A model linking the functions of TOR complexes and Paf1C during quiescence
The observed reduction in TORC2 activeness could explain the reduced H3K9me2 levels in heterochromatic regions in G0 cells. TORC2 was recently shown to be required for subtelomeric silencing and heterochromatin assembly in vegetative cells [xiii]. The Gad8 kinase is nowadays in the nucleus [19]; therefore, information technology is therefore conceivable that the TORC2/Gad8 pathway directly regulates Paf1C during quiescence. Nosotros showed that Torin1 treatment de-repressed some subtelomeric genes simply did not impact leo1∆ cells (Fig. 6a, b). Torin1 inhibits both the TORC1 and TORC2 complexes in fission yeast [20]. Therefore, both TOR complexes possibly act upstream of Paf1C in the control of heterochromatin dynamics in quiescent cells (Fig. 6c).
Conclusions
Here, nosotros demonstrated two of import concepts:
- 1.
During cellular quiescence in fission yeast, heterochromatic regions become very dynamic, especially in subtelomeres.
- 2.
Paf1C, acting downstream of TOR complexes, is a key player in the dynamic regulation of heterochromatin in quiescent cells, allowing changes in cistron expression and adaptation to the new cellular environs.
Paf1C is conserved in eukaryotes [7], and a component of Paf1C was recently implicated in a jail cell differentiation procedure in Drosophila [viii]. The dynamic regulation of heterochromatin in quiescent cells by Paf1C may also contribute to the office of Paf1C in cell differentiation in multicellular eukaryotes.
Methods
Yeast strains and growth weather
A selection of strains from the Bioneer library version 5 carrying deletions of genes encoding various chromatin regulators was backcrossed, mat1-Chiliad smt-0 strains carrying cistron deletions marked by kanMX were selected, and the mating blazon was confirmed past PCR [21]. The following strains were used for farther studies: PB1623 mat1-M smt-0; PB2423 mat1-M smt-0 leo1∆; PB2426 mat1-M smt-0 hht1∆; and PB2428 mat1-Thou smt-0 clr4∆. For liquid cultures, pombe minimal glutamate (PMG) and PMG without nitrogen (PMG-North) media were used. For the genetic screen, survival during long-term quiescence was followed in xx-ml PMG-Northward cultures at 32 °C for 48 days by plating aliquots of cells on rich medium (yeast excerpt with supplements, Aye) at regular intervals.
Quiescence induction
Cells were grown in PMG medium at xxx °C to a density of OD600 ~ 0.25. Quiescence was induced by washing cells twice with PMG-N medium and resuspending them in an equal volume of PMG-N medium.
Propidium iodide (PI) staining and flow cytometry
Cells grown in PMG or PMG-N medium were collected and stock-still with seventy% ethanol. So, cells were washed with 50 mM sodium citrate (pH 7.0) and treated with 240 µg/ml RNase A in 50 mM sodium citrate (pH 7.0) for one h at 37 °C. Cells were stained with ane.25 µg/ml PI in 50 mM sodium citrate (pH 7.0). Stained cells were imaged with a confocal microscope using the Cy5 channel. The Dna content was monitored using a FACSCalibur (Becton Dickinson). Information analysis was carried out with Prison cell Quest software.
Viability assay
Viability assays were performed according to the protocol described in Joh et al. [6], with pocket-sized modifications. A i ml aliquot of cell suspensions was taken from incubated cell cultures at the indicated time points and stained with Phloxin B (5 µg/ml final concentration) for 15 min. Cells were washed with one × phosphate-buffered saline (PBS) and imaged with an Axioplan ii fluorescence microscope and ImageJ software using the FITC and DIC channels. A minimum of 400 cells was counted for each measurement using ImageJ software, and survival rates were calculated based on the number of dead (stained and greenish) cells divided past the total number of cells (green and stain-gratuitous) counted.
Chromatin immunoprecipitation (Fleck) analysis
Chromatin was extracted from cultures in duplicate and subjected to immunoprecipitation. Samples were immediately subjected to formaldehyde cross-linking (final concentration one%), and chromatin was isolated as described by Durand-Dubief et al [22]. ChIP was performed using 4 µg of anti-H3K9me2 antibodies (Abcam, ab1220) per 30 µl of chromatin excerpt. DNA was recovered with QIAquick PCR Purification columns (Qiagen).
ChIP microarray
For microarray hybridization, immunoprecipitated Deoxyribonucleic acid was amplified as described by Durand-Dubief et al [22]. Fragmentation, labeling and hybridization to the Affymetrix GeneChip S. pombe Tiling 1.0FR were performed by the BEA core facility at Novum (http://www.bea.ki.se) according to Affymetrix standard protocols. For domainogram visualization, raw data from Affymetrix (.CEL format) were normalized using Affymetrix Tiling Assay Software (TAS) v1.1. Duplicate data were normalized to input using quantile normalization plus scaling using a bandwidth of 100. Data obtained from TAS were then directly loaded into the Seqmonk program, and the antibody background was removed using data for the clr4∆ mutant. Data were visualized using the following parameters: number of classes, 30; min probe count, 30; and max probe count, 100. Raw and normalized Scrap microarray information have been submitted to the Gene Expression Motorbus under accession number GSE116038.
RNA isolation
Cells were harvested by centrifugation at 3000 rpm and 25 °C. Then, cells were washed in one case with PBS (pH 7.four). The pellet was re-suspended in 500 µl of RNA extraction buffer (two% Triton X-100, 1% SDS, 100 mM NaCl, 10 mM Tris–HCl (pH 8.0), and 1 mM EDTA (pH 8.0)) and 500 µl of acrid phenol (pH 5.2). Then, 500 µl of acid-washed glass beads were added to the tubes. The tubes were vortexed vigorously at 4 °C to disrupt the cells. The tubes were centrifuged at 13,000 rpm for 5 min at 4 °C, and the upper aqueous phase was collected into a gel stage tube containing an equal volume of chloroform. After mixing, the tubes were centrifuged at 13,000 rpm for v min at 25 °C, and the upper aqueous phase was collected. Ammonium acetate (pH five.2) was added to a final concentration of ii.5 K, along with ii.5 volumes of ethanol. The tubes were stored at − 20 °C overnight for RNA precipitation. The precipitated RNA was done one time with 70% ethanol and dissolved in RNase-free h2o. DNase I treatment was performed using TURBO DNase (Thermo Fisher Sci.) post-obit the manufacturer'southward instructions.
RNA-seq
To remove rRNA, 4 µg of purified total RNA was treated with a ScriptSeq Complete Gold Kit (Yeast) (Epicentre). A full of xl ng of rRNA-depleted stocks were used to generate sequencing libraries using a ScriptSeq v2 RNA-seq Library Preparation Kit (Epicentre). Samples were quantified using a Qubit (HS dsDNA) and sequenced using an Illumina HiSeq 2000 platform (fifty cycles, single-terminate sequencing) at the BEA facility (Huddinge, Sweden) following the manufacturer's instructions. The raw HiSeq information (fastq files) were aligned to ASM294v2 using Bowtie2 with the default parameters. The ASM294v2.24 notation was downloaded from pombase.org. The aligned data (SAM files) were imported and normalized per meg reads. Data from independent biological duplicates were averaged. Identical reads were discarded to remove PCR artefacts. Signals were calculated as averages over 150 nt.
H3-rite
H3-RITE was carried out co-ordinate to the procedure described in [10] except for the changes in the growth media. Here, cells were cultured in PMG-Northward medium at xxx °C for 24 h afterwards the change from PMG medium. To induce the genetic switch, cells were treated with 1 µM β-oestradiol for iv h at 30 °C.
Torin1 treatment
Cells were grown in PMG-Due north medium at 28 °C to a density of OD600 ~ 0.25 and treated with Torin1 for 30 min at a final concentration of 25 µM or with an equivalent book of DMSO vehicle control [23].
RT-PCR
For RT-PCR analysis, full RNA was cleaned and treated with TURBO DNase (Thermo Fisher Scientific) according to the manufacturer's instructions. RT-PCR was performed using a SuperScript Iii First-Strand Synthesis System for RT-PCR (Thermo Fisher Scientific) according to the manufacturer's instructions. For RT-PCR, 400 ng of total RNA from S. pombe was used per reaction, and 50 ng of human RNA extracted from human bone osteosarcoma epithelial cells (U2OS cell line) was added as the spike-in control RNA for normalization of the expression levels.
qPCR
qPCR was performed using FastStart Universal SYBR Green Master (Rox) (Sigma-Aldrich) in a 7500 Fast Existent-Time PCR System (Practical Biosystems). For RT-qPCR, man GAPDH levels were measured for the normalization of RNA levels. The primer sequences used are shown in Table S1.
Western blotting
Protein was extracted from vegetative and quiescent cells. A total of twenty µg of poly peptide was separated past ten% SDS–Page and transferred to nitrocellulose membranes. Membranes were incubated with anti-phosphorylated Gad8 (custom-fabricated) (1:1000) and anti-actin (MP #691,001) (1:1000) antibodies, and immunoreactions were detected by an ECL (SuperSignal Detection System, Thermo Scientific) kit.
Availability of data and materials
Genome-broad information take been submitted to the Gene Expression Jitney under the accretion number GSE116038.
Abbreviations
- TOR:
-
target of rapamycin
- TORC2:
-
target of rapamycin complex 2
- Paf1C RNA:
-
polymerase-associated factor 1 complex
- Agone:
-
argonaute
- H3-RITE:
-
recombination-induced tag exchange for histone H3
- PMG:
-
pombe minimal glutamate medium
References
-
Su SS, Tanaka Y, Samejima I, Tanaka K, Yanagida G. A nitrogen starvation-induced dormant G0 land in fission yeast: the establishment from uncommitted G1 state and its delay for return to proliferation. J Jail cell Sci. 1996;109(Pt 6):1347–57.
-
Sajiki K, Hatanaka M, Nakamura T, Takeda K, Shimanuki Grand, Yoshida T, et al. Genetic command of cellular quiescence in S. pombe. J Jail cell Sci. 2009;122:1418–29.
-
Takeda One thousand, Yanagida Thou. In quiescence of fission yeast, autophagy and the proteasome interact for mitochondrial maintenance and longevity. Autophagy. 2010;6:564–5.
-
Marguerat S, Schmidt A, Codlin S, Chen W, Aebersold R, Bahler J. Quantitative analysis of fission yeast transcriptomes and proteomes in proliferating and quiescent cells. Cell. 2012;151:671–83.
-
Roche B, Arcangioli B, Martienssen RA. RNA interference is essential for cellular quiescence. Science. 2016;354:11.
-
Joh RI, Khanduja JS, Calvo IA, Mistry Chiliad, Palmieri CM, Savol AJ, et al. Survival in quiescence requires the euchromatic deployment of Clr4/SUV39H by argonaute-associated small RNAs. Mol Cell. 2016;64:1088–101.
-
Jaehning JA. The Paf1 circuitous: platform or player in RNA polymerase II transcription? Biochim Biophys Acta. 2010;1799:379–88.
-
Bahrampour S, Thor S. Ctr9, a key component of the Paf1 circuitous, affects proliferation and last differentiation in the developing drosophila nervous system. G3 (Bethesda). 2016;6:3229–39.
-
Kowalik KM, Shimada Y, Flury V, Stadler MB, Batki J, Buhler M. The Paf1 complex represses small-RNA-mediated epigenetic cistron silencing. Nature. 2015;520:248–52.
-
Sadeghi L, Prasad P, Ekwall Yard, Cohen A, Svensson JP. The Paf1 circuitous factors Leo1 and Paf1 promote local histone turnover to modulate chromatin states in fission yeast. EMBO Rep. 2015;16:1673–87.
-
Cam HP, Sugiyama T, Chen ES, Chen X, FitzGerald PC, Grewal SI. Comprehensive assay of heterochromatin- and RNAi-mediated epigenetic command of the fission yeast genome. Nat Genet. 2005;37:809–19.
-
Yamanaka S, Mehta S, Reyes-Turcu Atomic number 26, Zhuang F, Fuchs RT, Rong Y, et al. RNAi triggered by specialized machinery silences developmental genes and retrotransposons. Nature. 2013;493:557–threescore.
-
Cohen A, Habib A, Laor D, Yadav Southward, Kupiec M, Weisman R. TOR circuitous 2 in fission yeast is required for chromatin-mediated gene silencing and assembly of heterochromatic domains at subtelomeres. J Biol Chem. 2018;293:8138–50.
-
Cohen A, Kupiec M, Weisman R. Glucose activates TORC2-Gad8 protein via positive regulation of the cAMP/army camp-dependent poly peptide kinase A (PKA) pathway and negative regulation of the Pmk1 protein-mitogen-activated protein kinase pathway. J Biol Chem. 2014;289:21727–37.
-
Hatano T, Morigasaki S, Tatebe H, Ikeda K, Shiozaki K. Fission yeast Ryh1 GTPase activates TOR complex 2 in response to glucose. Jail cell Cycle. 2015;14:848–56.
-
Matsuo T, Kubo Y, Watanabe Y, Yamamoto K. Schizosaccharomyces pombe AGC family kinase Gad8p forms a conserved signaling module with TOR and PDK1-similar kinases. EMBO J. 2003;22:3073–83.
-
Xu Y, Bernecky C, Lee CT, Maier KC, Schwalb B, Tegunov D, et al. Architecture of the RNA polymerase II-Paf1C-TFIIS transcription elongation complex. Nat Commun. 2017;8:15741.
-
Jih G, Iglesias N, Currie MA, Bhanu NV, Paulo JA, Gygi SP, et al. Unique roles for histone H3K9me states in RNAi and heritable silencing of transcription. Nature. 2017;547:463–7.
-
Cohen A, Kupiec M, Weisman R. Gad8 protein is found in the nucleus where it interacts with the MluI jail cell cycle box-bounden gene (MBF) transcriptional circuitous to regulate the response to Deoxyribonucleic acid replication stress. J Biol Chem. 2016;291:9371–81.
-
Matsuo T, Otsubo Y, Urano J, Tamanoi F, Yamamoto M. Loss of the TOR kinase Tor2 mimics nitrogen starvation and activates the sexual development pathway in fission yeast. Mol Cell Biol. 2007;27:3154–64.
-
Ekwall Thousand, Thon G. Mating-type determination in Schizosaccharomyces pombe. Cold Spring Harb Protoc. 2017;2017:pdb prot091728.
-
Durand-Dubief M, Ekwall Thousand. Chromatin immunoprecipitation using microarrays. Methods Mol Biol. 2009;529:279–95.
-
Atkin J, Halova 50, Ferguson J, Hitchin JR, Lichawska-Cieslar A, Jordan AM, et al. Torin1-mediated TOR kinase inhibition reduces Wee1 levels and advances mitotic commitment in fission yeast and HeLa cells. J Cell Sci. 2014;127:1346–56.
Acknowledgements
We thank the BEA facility, peculiarly T. Damdimopoulos, for help in processing cistron expression information.
Funding
Piece of work in the K.E. laboratory was supported by grants from the Swedish Cancer Club (C.F.) and the Swedish Inquiry Quango (V.R.). Thou.East. was the recipient of a Wenner-Gren stipend for a breather stay at the Pasteur Institute, France. E.O. was the recipient of a postdoctoral fellowship and travel support from the JSPS Program for Advancing Strategic International Networks to Advance the Circulation of Talented Researchers (S2704). Funding was provided by Cancerfonden, Vetenskapsrådet, Wenner-Gren Foundation and Japan Society for the Promotion of Science.
Writer data
Affiliations
Contributions
EO performed most of the experiments and wrote the showtime typhoon of the manuscript. KE planned and supervised the study and helped write the manuscript. MDD analyzed the genome-wide Fleck data. AC and RW contributed to experiments on Gad8-P. CS performed strain construction, and BA helped program the experiments. VM helped with experiments during the revisions. J-IN gave valuable advice and helped with the supervision of EO. All authors commented on and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ideals declarations
Ethical approving
Not applicative for research on fission yeast.
Consent for publication
All authors concur to the publication of this manuscript.
Competing interests
The authors declare that they take no competing interests.
Boosted information
Publisher'south Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional files
Rights and permissions
Open Admission This commodity is distributed under the terms of the Artistic Commons Attribution iv.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you lot requite advisable credit to the original writer(s) and the source, provide a link to the Creative Eatables license, and point if changes were made. The Artistic Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/ane.0/) applies to the data made available in this commodity, unless otherwise stated.
Reprints and Permissions
Virtually this article
Cite this article
Oya, Due east., Durand-Dubief, Grand., Cohen, A. et al. Leo1 is essential for the dynamic regulation of heterochromatin and factor expression during cellular quiescence. Epigenetics & Chromatin 12, 45 (2019). https://doi.org/10.1186/s13072-019-0292-7
-
Received:
-
Accepted:
-
Published:
-
DOI : https://doi.org/10.1186/s13072-019-0292-7
Keywords
- Paf1C
- Cellular quiescence
- Heterochromatin
- Gene expression
- Fission yeast
Source: https://epigeneticsandchromatin.biomedcentral.com/articles/10.1186/s13072-019-0292-7
Posted by: padgettmilesse.blogspot.com


0 Response to "Which Of The Following Chromatin Changes Will Lead To Upregulation Of Expression?"
Post a Comment